While still emerging in clinical practice, HLA-restricted therapies are gaining traction in clinical research and early-stage precision medicine applications (see our relevant blog for further details). As these HLA-restricted therapies often specifically benefit patients carrying a specific HLA allele, HLA testing is becoming an increasingly important step to match the right patient to the right (HLA-restricted) treatment. GenDx offers both Sanger Sequencing-based and Next-Generation Sequencing (NGS)-based solutions for HLA typing. While Sanger sequencing has long been a trusted method for HLA typing, NGS-based HLA typing provides key benefits that make it well-suited for evolving high-throughput and high-resolution diagnostic needs for molecular (companion) diagnostic solutions. In this overview, we highlight key concepts and introduce several technical considerations relevant to the sequencing technology used for HLA typing. This overview reflects current technological perspectives, including solutions developed by GenDx.
Sanger sequencing relies on chain-terminating fluorescent nucleotides to determine DNA sequence. NGS encompasses a range of technologies with different detection principles, including sequencing-by-synthesis with reversible terminators, semiconductor-based detection, and nanopore-based signal measurement. However, the key difference lies in their sequencing capacity. While Sanger sequences one DNA fragment at a time, with NGS it is possible to sequence millions of fragments at the same time. This approach greatly increases throughput, resolution and the ability to detect novel or rare variants with deep sequencing. Additionally, NGS enables scalable whole-gene coverage depending on amplification strategy and assay design, whereas Sanger sequencing is typically applied to targeted regions, such as exons. While both Sanger and NGS methods offer high accuracy, the errors associated with these methods differ. Sanger sequencing shows very high per-read accuracy and NGS shows high consensus accuracy through deep sequencing coverage that allows for error correction.
While performance may vary depending on assay design, laboratory workflow, and clinical context, employing NGS-based HLA typing can provide many benefits over Sanger-based HLA typing, as summarized in the table below. Note that the information provided below is tailored to short-read NGS using sequencing-by-synthesis technology.
Comparison of NGS-based and Sanger-based typing for Class I HLA genes.
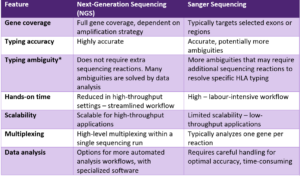
Despite several clear advantages, NGS-based HLA typing may also come with challenges, including variable coverage across genomic regions, potential amplification bias, and the need for complex bioinformatics pipelines. GenDx has addressed these challenges and offers NGS-based whole gene solutions for the HLA Class I genes (HLA-A, HLA-B and HLA-C) and for several Class II genes (HLA-DRB1, HLA-DQB1, HLA-DPB1, HLA-DRB3/4/5, HLA-DQA1 and HLA-DPA1), including GenDx’s HLA typing software NGSengine.
Difference between sequencing techniques in resolving typing ambiguities
For accurate HLA analysis, it is important to resolve as many typing ambiguities as possible. Ambiguities refer to the situation in which a typing result cannot uniquely identify the specific HLA alleles present in a sample, resulting in multiple possible allele or genotype combinations that are consistent with the data. While Sanger-based HLA typing may offer high accuracy for targeted regions, there may be a need for additional sequencing to resolve HLA typing ambiguities. NGS-based HLA typing offers a significant advantage, as many ambiguities can be resolved through data analysis. Nevertheless, some ambiguities may still require additional investigation.
When it comes to obtaining unambiguous typing results, it is relevant to:
(1) sequence as much of the gene as possible and,
(2) have “phasing” information over the full amplicon.
Resolving the exact cis/trans position of heterozygous positions is enabled by sequencing reads that span multiple heterozygous positions, combined with advanced software algorithms. This process is called “phasing”. The possibility to phase heterozygous positions depends on read length, insert size, and assay design and may still be limited by factors such as coverage variability and complex genomic regions.
Sanger and NGS show important differences in how and how well the ambiguities can be resolved. Figure 1 illustrates how different sequencing approaches impact ambiguity resolution.
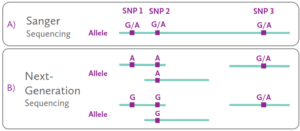
Figure 1. Schematic visualization how different sequencing technologies resolve HLA alleles.
A) When using Sanger-based sequencing, short DNA sequences are obtained containing information of both alleles in one trace. The use of group-specific sequencing primers (GSSPs) is necessary to separate and identify the two alleles. Moreover, due to the inability to link distant variants within a single read, it is not always possible to know the cis/trans position of distant heterozygous positions. As such, typing ambiguities that cannot be resolved are common.
B) When using NGS-based HLA typing, amplicon DNA is randomly fragmented into smaller fragments before sequencing. Heterozygous positions can be phased when the distance between the two heterozygous positions is not too large (left side of the schematic). In case the distance is too large, phasing cannot be achieved (right side of the schematic). Paired-end sequencing increases effective phasing distance, as it effectively links sequencing data across a span much larger than the read length itself.
Research Use of NGS-based HLA typing data
A high-resolution, whole-gene approach facilitated by NGS allows for deep investigation of a biomarker. Many of the current CDx applications rely on lower-resolution HLA typing as treatment eligibility is typically determined by second-field (protein-level) HLA typing resolution, although requirements may vary depending on the therapy and clinical context. Emerging applications may benefit from higher-resolution and/or whole-gene data. A whole-gene NGS approach generates data that allows for high-resolution analysis up to the fourth field, which is currently mainly relevant for research applications. Using this high-resolution information for research purposes can potentially provide interesting insights when stratifying non-responders, low-responders and responders and allows for exploring relationships between genetic variation and treatment response. Combined with powerful multiplexing capabilities, NGS opens the door to both retrospective studies and efficient pre-screening of patients during clinical investigations. Increasing the resolution of HLA typing in precision medicine, in the context of routine HLA typing and HLA screening for eligibility purposes, may contribute to more refined patient stratification and could help improve understanding of treatment response variability.
Interested in learning more?
Do you want to know more about HLA-restricted therapies, the importance of HLA in the Precision Medicine field or the impact of HLA on therapy success? Check out our previous blog posts and stay tuned for the next one!
Do you want to learn more about our Companion Diagnostics services and how we can be your partner from start to finish? Read more on our dedicated Companion Diagnostics webpage or get in touch with our team.
About the authors
Jules Petit – Project manager CDx

Jules holds a PhD in Molecular Immunology. After obtaining his PhD, Jules worked in different positions, both in an academic and in a pharmacological industry setting, before starting at GenDx. At GenDx he manages the different Companion Diagnostics programs and supports the scientific, strategic and regulatory integration of these projects.
Eleni Draina – Product Manager

Eleni holds a Master’s degree in Molecular Biology and currently works as a Product Manager at GenDx. In this role, she is responsible for a diverse portfolio of products—including CDx—and ensures that all customer-facing documentation is accurate, accessible, and up to date.
Relevant references:
Gormally, M. V., Chen, M. F., Noronha, A. M., Panageas, K., Reynolds, M., Kohlasch, K., … & Donoghue, M. T. (2025). Next-generation sequencing for HLA genotype screening and matching to HLA-restricted therapies. JAMA oncology, 11(1), 74-76.
Rozemuller, E. H., & Mulder, W. (2025). High‐Resolution HLA Typing: Sanger Sequencing, NGS, Long‐range Sequencing. Immunogenetics and Histocompatibility: A Guide for Practitioners, 413-441.
Bravo-Egana, V., Sanders, H., & Chitnis, N. (2021). New challenges, new opportunities: next generation sequencing and its place in the advancement of HLA typing. Human Immunology, 82(7), 478-487.
Cornaby, C., & Weimer, E. T. (2020). HLA Typing by Next-Generation Sequencing: Lessons Learned and Future Applications. Advances in Molecular Pathology, 3, 199-205.
Geo JA, Ameen R, Al Shemmari S, Thomas J. Advancements in HLA Typing Techniques and Their Impact on Transplantation Medicine. Med Princ Pract. 2024;33(3):215-231. doi: 10.1159/000538176. Epub 2024 Mar 5. PMID: 38442703; PMCID: PMC11175610.


