In the world of transplantation and clinical immunology, HLA (Human Leukocyte Antigen) typing is the ultimate “barcode” of human identity. It is the system our bodies use to differentiate “self” from “non-self.” Because this system is the most diverse part of the human genome, we need a naming convention that is as precise as the DNA itself.
With thousands of new alleles discovered every year, scientists rely on a standardized nomenclature, a biological “IP address” that allows clinicians and researchers worldwide to speak the exact same language.
The anatomy of an allele name
The length of an allele’s name is determined by its unique sequence and how closely it resembles its nearest known relative. While every allele is assigned at least a four-digit designation (covering the first two fields), additional digits are added when further genetic distinction is required.
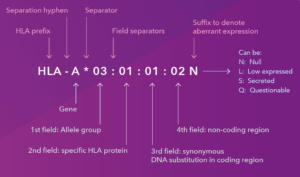
At first glance, an HLA designation like HLA-DRB1*13:01:01:02N looks like a complex string of random characters. In reality, it is a highly structured map.
Here is how to read it:
1. The prefix and gene (HLA-DRB1)
Before the asterisk, we identify the specific “location” the gene lives in.
-
HLA: The region.
-
DRB1: The specific locus (location) on the chromosome.
2. The first field: The type (*13)
The digits before the first colon usually describe the serological type. This is a callback to the old days* of “low resolution” typing. If you were an HLA specialist in the 1980s, you would recognize “13” as the DR13 antigen. It’s the broad family name.
3. The second field: The subtype (:01)
Now we get into the proteins. If two alleles have different numbers in this field (e.g., :01 vs :02), it means there is a change in the amino acid sequence. For the specialist, this is the most critical field for transplantation, as changes here can alter the shape of the peptide-binding groove, potentially triggering an immune response.
4. The third field: Silent substitutions (:01)
Sometimes, DNA changes but the protein stays exactly the same. These are called synonymous or “silent” mutations.
-
Why it matters: Even if the protein doesn’t change, these variations help researchers track human migration and genetic evolution.
5. The fourth field: The non-coding regions (:02)
This field looks at the “dark matter” of the gene, the introns or the flanking regions (5’ and 3’ UTRs). These are areas of the DNA that aren’t turned into protein but might affect how much of the protein is made.
The “special suffixes”: Nature’s plot twists
Sometimes, a gene is present but it doesn’t behave. This is where the letter at the end comes in. Think of these as the “status updates” of the HLA world:
-
N (Null): The “Ghost” allele. The DNA is there, but no protein is expressed. In a transplant, an “N” allele is effectively invisible to the immune system.
-
L (Low): The “Quiet” allele. The protein reaches the cell surface, but in significantly lower amounts than normal.
-
S (Secreted): The “Rule-Breaker.” Instead of sitting on the cell surface like a flag, this protein is shed and found floating freely in fluids.
-
Q (Questionable): The “Mystery.” A mutation exists that usually messes up expression in other genes, but we haven’t confirmed exactly what it’s doing here yet.
*Why not just use old names?
Before 2010, the naming system didn’t use colons, which led to a “naming ceiling.” As we discovered more than 99 subtypes for a single group, we ran out of numbers! The modern colon-delimited system provides an infinite runway for future discoveries.
Want to dive deeper into the latest allele updates? Check out the IPD-IMGT/HLA Database for a full list of the thousands of alleles currently recognized by the World Health Organization (WHO) Nomenclature Committee.


